- Mahotas - Home
- Mahotas - Introduction
- Mahotas - Computer Vision
- Mahotas - History
- Mahotas - Features
- Mahotas - Installation
- Mahotas Handling Images
- Mahotas - Handling Images
- Mahotas - Loading an Image
- Mahotas - Loading Image as Grey
- Mahotas - Displaying an Image
- Mahotas - Displaying Shape of an Image
- Mahotas - Saving an Image
- Mahotas - Centre of Mass of an Image
- Mahotas - Convolution of Image
- Mahotas - Creating RGB Image
- Mahotas - Euler Number of an Image
- Mahotas - Fraction of Zeros in an Image
- Mahotas - Getting Image Moments
- Mahotas - Local Maxima in an Image
- Mahotas - Image Ellipse Axes
- Mahotas - Image Stretch RGB
- Mahotas Color-Space Conversion
- Mahotas - Color-Space Conversion
- Mahotas - RGB to Gray Conversion
- Mahotas - RGB to LAB Conversion
- Mahotas - RGB to Sepia
- Mahotas - RGB to XYZ Conversion
- Mahotas - XYZ to LAB Conversion
- Mahotas - XYZ to RGB Conversion
- Mahotas - Increase Gamma Correction
- Mahotas - Stretching Gamma Correction
- Mahotas Labeled Image Functions
- Mahotas - Labeled Image Functions
- Mahotas - Labeling Images
- Mahotas - Filtering Regions
- Mahotas - Border Pixels
- Mahotas - Morphological Operations
- Mahotas - Morphological Operators
- Mahotas - Finding Image Mean
- Mahotas - Cropping an Image
- Mahotas - Eccentricity of an Image
- Mahotas - Overlaying Image
- Mahotas - Roundness of Image
- Mahotas - Resizing an Image
- Mahotas - Histogram of Image
- Mahotas - Dilating an Image
- Mahotas - Eroding Image
- Mahotas - Watershed
- Mahotas - Opening Process on Image
- Mahotas - Closing Process on Image
- Mahotas - Closing Holes in an Image
- Mahotas - Conditional Dilating Image
- Mahotas - Conditional Eroding Image
- Mahotas - Conditional Watershed of Image
- Mahotas - Local Minima in Image
- Mahotas - Regional Maxima of Image
- Mahotas - Regional Minima of Image
- Mahotas - Advanced Concepts
- Mahotas - Image Thresholding
- Mahotas - Setting Threshold
- Mahotas - Soft Threshold
- Mahotas - Bernsen Local Thresholding
- Mahotas - Wavelet Transforms
- Making Image Wavelet Center
- Mahotas - Distance Transform
- Mahotas - Polygon Utilities
- Mahotas - Local Binary Patterns
- Threshold Adjacency Statistics
- Mahotas - Haralic Features
- Weight of Labeled Region
- Mahotas - Zernike Features
- Mahotas - Zernike Moments
- Mahotas - Rank Filter
- Mahotas - 2D Laplacian Filter
- Mahotas - Majority Filter
- Mahotas - Mean Filter
- Mahotas - Median Filter
- Mahotas - Otsu's Method
- Mahotas - Gaussian Filtering
- Mahotas - Hit & Miss Transform
- Mahotas - Labeled Max Array
- Mahotas - Mean Value of Image
- Mahotas - SURF Dense Points
- Mahotas - SURF Integral
- Mahotas - Haar Transform
- Highlighting Image Maxima
- Computing Linear Binary Patterns
- Getting Border of Labels
- Reversing Haar Transform
- Riddler-Calvard Method
- Sizes of Labelled Region
- Mahotas - Template Matching
- Speeded-Up Robust Features
- Removing Bordered Labelled
- Mahotas - Daubechies Wavelet
- Mahotas - Sobel Edge Detection
Mahotas - Reversing Haar Transform
Reversing Haar transform refers to the technique of reconstructing the original image from a Haar transformed image. Before understanding reverse Haar transformation, let us learn about Haar transformation.
Haar transformation is a technique that converts an image from pixel intensity values to wavelet coefficients (values that represent different frequencies of an image).
In Haar transformation, the image is broken into a set of orthonormal basis functions called Haar wavelets.
Reversing Haar transformation converts the wavelet coefficients into pixel intensity values by combining the Haar wavelets in a specific manner (as discussed below).
Reversing Haar Transform in Mahotas
In Mahotas, we can perform reverse Haar transformation by using the mahotas.ihaar() function. Following is the basic approach to perform the inverse Haar transformation −
First, obtain the Haar wavelet coefficients from Haar transformation.
Next, multiply each coefficient by a scaling factor and the Haar wavelet. For the Haar wavelet, the scaling factor is usually $\mathrm{1/\sqrt{2}}$ for the approximation coefficients and 1 for the detail coefficients.
Then, sum up these scaled coefficients for both the high frequency (detail) and the low frequency (approximation) coefficients.
Finally, combine the reconstructed coefficients and perform normalization if the pixel values are not within the range of 0 to 255.
Once these steps are completed, the original image is reconstructed from the Haar transformed image.
The mahotas.ihaar() function
The mahotas.ihaar() function takes a Haar transformed image as an input and returns the original grayscale image as output.
The reverse image is a perfect reconstruction of the original image since Haar transformation is a reversible process.
Syntax
Following is the basic syntax of the ihaar() function in mahotas −
mahotas.ihaar(f, preserve_energy=True, inline=False)
Where,
f − It is the input image.
preserve_energy (optional) − It specifies whether to preserve the energy of the output image (default is True).
inline (optional) − It specifies whether to return a new image or modify the input image (default is False).
Example
In the following example, we are using the mh.ihaar() function to reverse the effect of Haar transformation on an image.
import mahotas as mh
import numpy as np
import matplotlib.pyplot as mtplt
# Loading the image
image = mh.imread('sun.png')
# Converting it to grayscale
image = mh.colors.rgb2gray(image)
# Applying Haar transformation
haar_transform = mh.haar(image)
# Reversing Haar transformation
reverse_haar = mh.ihaar(haar_transform)
# Creating a figure and axes for subplots
fig, axes = mtplt.subplots(1, 3)
# Displaying the original image
axes[0].imshow(image, cmap='gray')
axes[0].set_title('Original Image')
axes[0].set_axis_off()
# Displaying the Haar transformed image
axes[1].imshow(haar_transform, cmap='gray')
axes[1].set_title('Haar Transformed Image')
axes[1].set_axis_off()
# Displaying the reversed image
axes[2].imshow(reverse_haar, cmap='gray')
axes[2].set_title('Reverse Haar Image')
axes[2].set_axis_off()
# Adjusting spacing between subplots
mtplt.tight_layout()
# Showing the figures
mtplt.show()
Output
Following is the output of the above code −

Without Preserving Energy
We can also reverse the effects of Haar transformation on an image without preserving its energy. The energy of the image refers to its brightness and it can change when an image is being transformed.
In mahotas, we can set the preserve_energy parameter to 'False' in the mh.ihaar() function to prevent the energy preservation. Hence, the brightness of the output image will be different from the original input image.
If this parameter is set to True, then the output image and the input image will have the same brightness.
Example
In the example mentioned below, we are performing reverse Haar transformation on an image without preserving its energy.
import mahotas as mh
import numpy as np
import matplotlib.pyplot as mtplt
# Loading the image
image = mh.imread('tree.tiff')
# Converting it to grayscale
image = mh.colors.rgb2gray(image)
# Applying Haar transformation
haar_transform = mh.haar(image)
# Reversing Haar transformation
reverse_haar = mh.ihaar(haar_transform, preserve_energy=False)
# Creating a figure and axes for subplots
fig, axes = mtplt.subplots(1, 3)
# Displaying the original image
axes[0].imshow(image, cmap='gray')
axes[0].set_title('Original Image')
axes[0].set_axis_off()
# Displaying the Haar transformed image
axes[1].imshow(haar_transform, cmap='gray')
axes[1].set_title('Haar Transformed Image')
axes[1].set_axis_off()
# Displaying the reversed image
axes[2].imshow(reverse_haar, cmap='gray')
axes[2].set_title('Reverse Haar Image')
axes[2].set_axis_off()
# Adjusting spacing between subplots
mtplt.tight_layout()
# Showing the figures
mtplt.show()
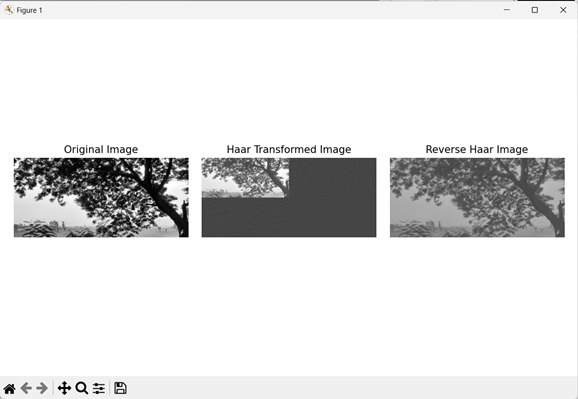
Output
Output of the above code is as follows −
Inline Reverse Haar Transformation
Another way we can reverse Haar transformation is by performing inline reverse Haar transformation. Inline refers to applying the transformation on the original image itself without creating a new image, thus saving space during transformation.
In mahotas, inline reverse Haar transformation can be achieved by setting the inline parameter to the boolean value 'True' in the mh.ihaar() function.
Example
Here, we are performing inline reverse Haar transformation on a Haar transformed image.
import mahotas as mh
import numpy as np
import matplotlib.pyplot as mtplt
# Loading the image
image = mh.imread('sea.bmp')
# Converting it to grayscale
image = mh.colors.rgb2gray(image)
# Reversing Haar transformation
mh.ihaar(mh.haar(image), inline=True)
# Creating a figure and axes for subplots
fig, axes = mtplt.subplots(1, 1)
# Displaying the reversed image
axes.imshow(image, cmap='gray')
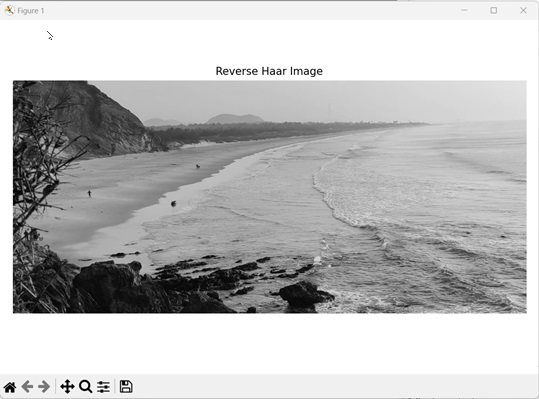
axes.set_title('Reverse Haar Image')
axes.set_axis_off()
# Adjusting spacing between subplots
mtplt.tight_layout()
# Showing the figures
mtplt.show()
Output
After executing the above code, we get the following output −