- Mahotas - Home
- Mahotas - Introduction
- Mahotas - Computer Vision
- Mahotas - History
- Mahotas - Features
- Mahotas - Installation
- Mahotas Handling Images
- Mahotas - Handling Images
- Mahotas - Loading an Image
- Mahotas - Loading Image as Grey
- Mahotas - Displaying an Image
- Mahotas - Displaying Shape of an Image
- Mahotas - Saving an Image
- Mahotas - Centre of Mass of an Image
- Mahotas - Convolution of Image
- Mahotas - Creating RGB Image
- Mahotas - Euler Number of an Image
- Mahotas - Fraction of Zeros in an Image
- Mahotas - Getting Image Moments
- Mahotas - Local Maxima in an Image
- Mahotas - Image Ellipse Axes
- Mahotas - Image Stretch RGB
- Mahotas Color-Space Conversion
- Mahotas - Color-Space Conversion
- Mahotas - RGB to Gray Conversion
- Mahotas - RGB to LAB Conversion
- Mahotas - RGB to Sepia
- Mahotas - RGB to XYZ Conversion
- Mahotas - XYZ to LAB Conversion
- Mahotas - XYZ to RGB Conversion
- Mahotas - Increase Gamma Correction
- Mahotas - Stretching Gamma Correction
- Mahotas Labeled Image Functions
- Mahotas - Labeled Image Functions
- Mahotas - Labeling Images
- Mahotas - Filtering Regions
- Mahotas - Border Pixels
- Mahotas - Morphological Operations
- Mahotas - Morphological Operators
- Mahotas - Finding Image Mean
- Mahotas - Cropping an Image
- Mahotas - Eccentricity of an Image
- Mahotas - Overlaying Image
- Mahotas - Roundness of Image
- Mahotas - Resizing an Image
- Mahotas - Histogram of Image
- Mahotas - Dilating an Image
- Mahotas - Eroding Image
- Mahotas - Watershed
- Mahotas - Opening Process on Image
- Mahotas - Closing Process on Image
- Mahotas - Closing Holes in an Image
- Mahotas - Conditional Dilating Image
- Mahotas - Conditional Eroding Image
- Mahotas - Conditional Watershed of Image
- Mahotas - Local Minima in Image
- Mahotas - Regional Maxima of Image
- Mahotas - Regional Minima of Image
- Mahotas - Advanced Concepts
- Mahotas - Image Thresholding
- Mahotas - Setting Threshold
- Mahotas - Soft Threshold
- Mahotas - Bernsen Local Thresholding
- Mahotas - Wavelet Transforms
- Making Image Wavelet Center
- Mahotas - Distance Transform
- Mahotas - Polygon Utilities
- Mahotas - Local Binary Patterns
- Threshold Adjacency Statistics
- Mahotas - Haralic Features
- Weight of Labeled Region
- Mahotas - Zernike Features
- Mahotas - Zernike Moments
- Mahotas - Rank Filter
- Mahotas - 2D Laplacian Filter
- Mahotas - Majority Filter
- Mahotas - Mean Filter
- Mahotas - Median Filter
- Mahotas - Otsu's Method
- Mahotas - Gaussian Filtering
- Mahotas - Hit & Miss Transform
- Mahotas - Labeled Max Array
- Mahotas - Mean Value of Image
- Mahotas - SURF Dense Points
- Mahotas - SURF Integral
- Mahotas - Haar Transform
- Highlighting Image Maxima
- Computing Linear Binary Patterns
- Getting Border of Labels
- Reversing Haar Transform
- Riddler-Calvard Method
- Sizes of Labelled Region
- Mahotas - Template Matching
- Speeded-Up Robust Features
- Removing Bordered Labelled
- Mahotas - Daubechies Wavelet
- Mahotas - Sobel Edge Detection
Mahotas - Filtering Regions
Filtering regions refers to excluding specific regions of a labeled image based on certain criteria. A commonly used criteria for filtering regions is based on their size. By specifying a size limit, regions that are either too small or too large can be excluded to get a clean output image.
Another criterion for filtering regions is to check whether a region is bordered or not. By applying these filters, we can selectively remove or retain regions of interest in the image.
Filtering Regions in Mahotas
In Mahotas, we can convert filter regions of a labeled image by using the labeled.filter_labeled() function. This function applies filters to the selected regions of an image while leaving other regions unchanged.
Using the mahotas.labeled.filter_labeled() Function
The mahotas.labeled.filter_labeled() function takes a labeled image as input and removes unwanted regions based on certain properties. It identifies the regions based on the labels of an image.
The resultant image consists of only regions that match the filter criterion.
Syntax
Following is the basic syntax of the filter_labeled() function in mahotas −
mahotas.labeled.filter_labeled(labeled, remove_bordering=False, min_size=None, max_size=None)
where,
labeled − It is the array.
remove_bordering (optional) − It defines whether to remove regions touching the border.
min_size (optional) − It is the minimum size of the region that needs to be kept (default is no minimum).
max_size (optional) − It is the maximum size of the region that needs to be kept (default is no maximum).
Example
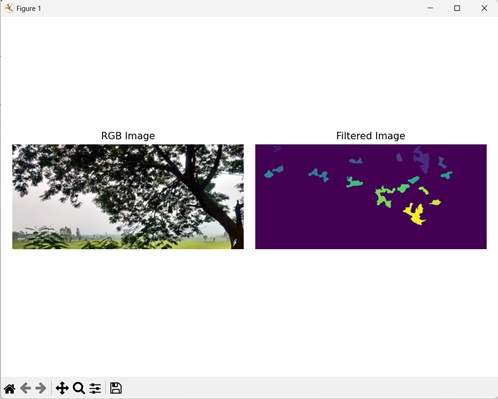
In the following example, we are filtering a labeled image to remove border pixels.
import mahotas as mh
import numpy as np
import matplotlib.pyplot as mtplt
# Loading the image
image_rgb = mh.imread('tree.tiff')
image = image_rgb[:,:,0]
# Applying gaussian filtering
image = mh.gaussian_filter(image, 4)
image = (image > image.mean())
# Converting it to a labeled image
labeled, num_objects = mh.label(image)
# Applying filters
filtered_image, num_objects = mh.labeled.filter_labeled(labeled,
remove_bordering=True)
# Creating a figure and axes for subplots
fig, axes = mtplt.subplots(1, 2)
# Displaying the original RGB image
axes[0].imshow(image_rgb)
axes[0].set_title('RGB Image')
axes[0].set_axis_off()
# Displaying the filtered image
axes[1].imshow(filtered_image)
axes[1].set_title('Filtered Image')
axes[1].set_axis_off()
# Adjusting spacing between subplots
mtplt.tight_layout()
# Showing the figures
mtplt.show()
Output
Following is the output of the above code −
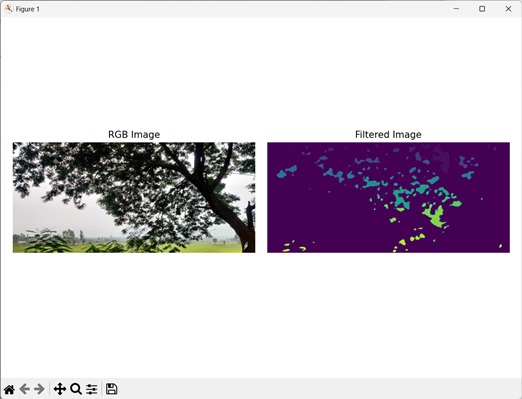
Filtering on Regions of Specific Size
We can also filter regions of specific size in an image. In this way, we can remove regions from labeled images which do not fall within a specific size limit (regions that are too small or too large).
In mahatos, we can achieve this by specifying values to the optional parameter min_size and max_size in the labeled.filter_label() function.
Example
The following example shows filtering a labeled image to remove regions of specific size.
import mahotas as mh
import numpy as np
import matplotlib.pyplot as mtplt
# Loading the image
image_rgb = mh.imread('tree.tiff')
image = image_rgb[:,:,0]
# Applying gaussian filtering
image = mh.gaussian_filter(image, 4)
image = (image > image.mean())
# Converting to a labeled image
labeled, num_objects = mh.label(image)
# Applying filters
filtered_image, num_objects = mh.labeled.filter_labeled(labeled, min_size=10,
max_size=50000)
# Create a figure and axes for subplots
fig, axes = mtplt.subplots(1, 2)
# Displaying the original RGB image
axes[0].imshow(image_rgb)
axes[0].set_title('RGB Image')
axes[0].set_axis_off()
# Displaying the filtered image
axes[1].imshow(filtered_image)
axes[1].set_title('Filtered Image')
axes[1].set_axis_off()
# Adjusting spacing between subplots
mtplt.tight_layout()
# Showing the figures
mtplt.show()
Output
Output of the above code is as follows −
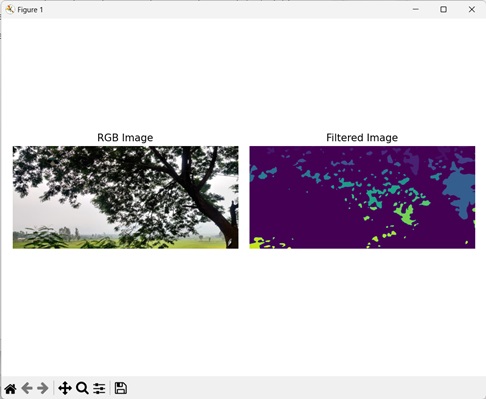
Filtering on Border Regions and Regions of Specific Size
We can filter bordered regions along with regions of a specific size in an image. In this, we remove regions which touch the border and regions which do not fall within a specific size limit.
In mahotas, we can do this by specifying values to the optional parameter min_size and max_size and setting the optional parameter remove_bordering to True in the labeled.filter_label() function.
Example
In this example, a filter is applied to remove border regions and regions of specific size of a labeled image.
import mahotas as mh
import numpy as np
import matplotlib.pyplot as mtplt
# Loading the image
image_rgb = mh.imread('tree.tiff')
image = image_rgb[:,:,0]
# Applying gaussian filtering
image = mh.gaussian_filter(image, 4)
image = (image > image.mean())
# Converting it to a labeled image
labeled, num_objects = mh.label(image)
# Applying filters
filtered_image, num_objects = mh.labeled.filter_labeled(labeled,
remove_bordering=True, min_size=1000, max_size=50000)
# Creating a figure and axes for subplots
fig, axes = mtplt.subplots(1, 2)
# Displaying the original RGB image
axes[0].imshow(image_rgb)
axes[0].set_title('RGB Image')
axes[0].set_axis_off()
# Displaying the filtered image
axes[1].imshow(filtered_image)
axes[1].set_title('Filtered Image')
axes[1].set_axis_off()
# Adjusting spacing between subplots
mtplt.tight_layout()
# Showing the figures
mtplt.show()
Output
The output produced is as shown below −